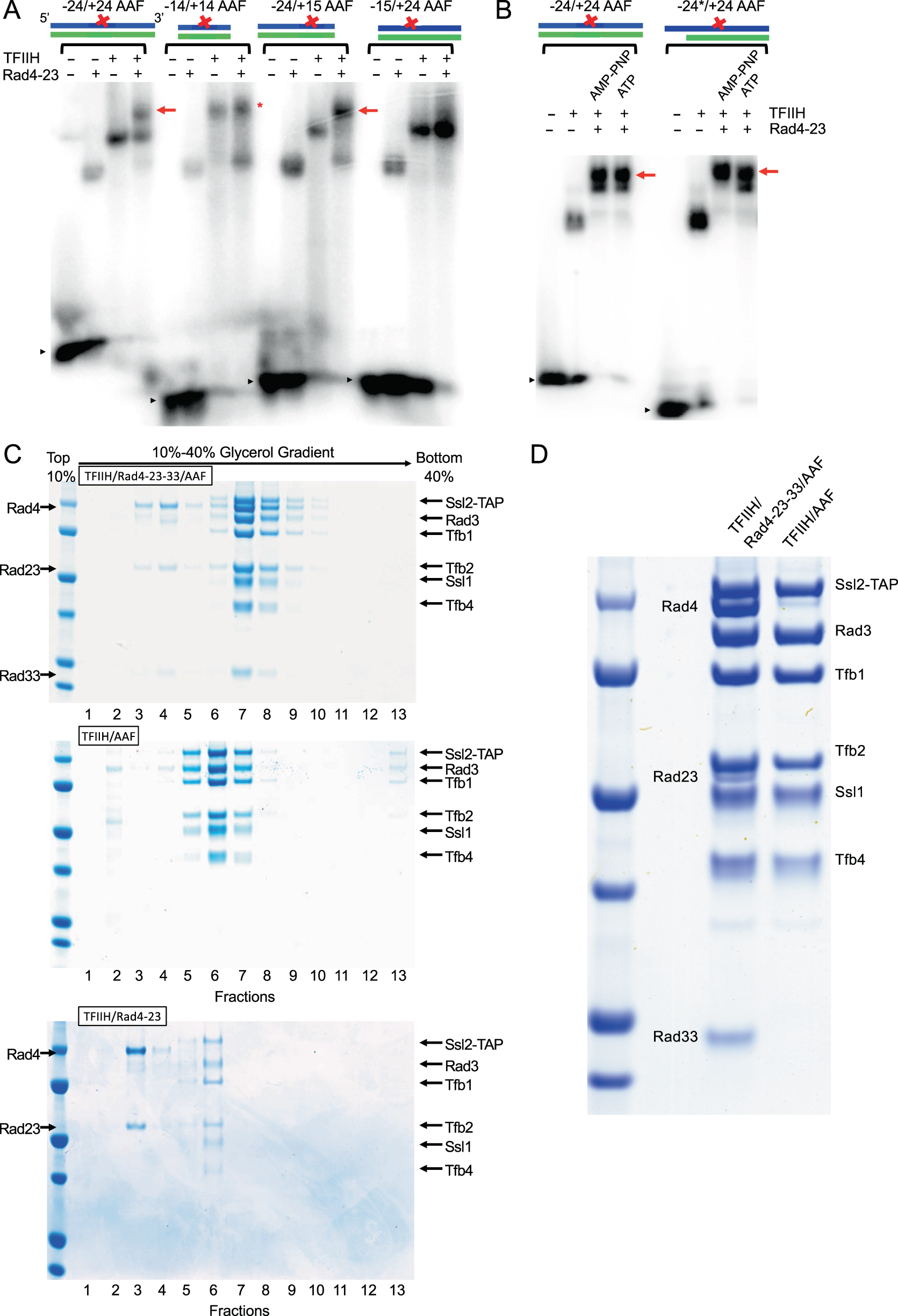
Cryo-EM structure of TFIIH/Rad4–Rad23–Rad33 in damaged DNA opening in nucleotide excision repair | Nature Communications
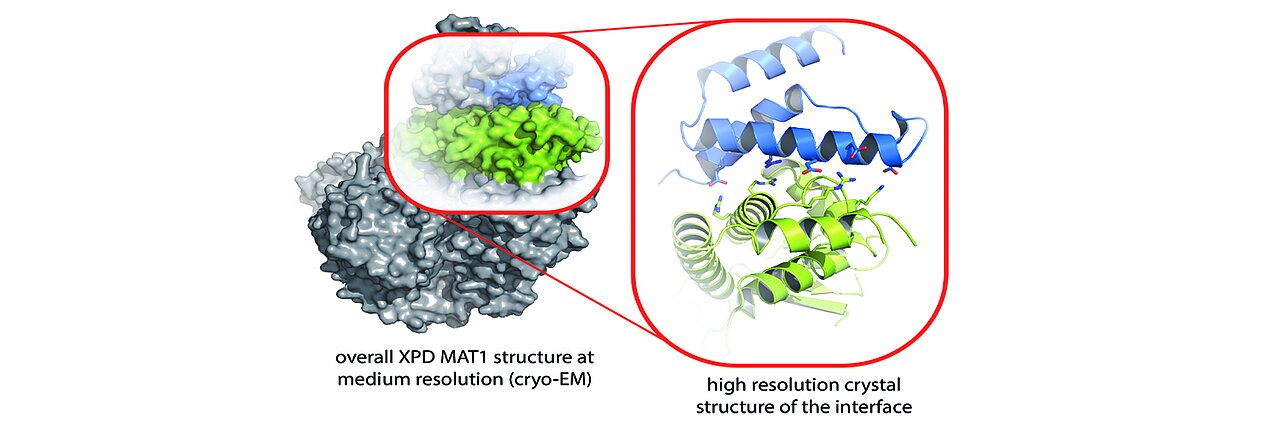
Switch for DNA repair tool discovered - Rudolf Virchow Center for Integrative and Translational Bioimaging

Jean-Marc EGLY | Institut de Génétique et de Biologie Moléculaire et Cellulaire, Strasbourg | IGBMC | Research profile

Phosphorylation of XPD drives its mitotic role independently of its DNA repair and transcription functions | Science Advances

Jean-Marc EGLY | Institut de Génétique et de Biologie Moléculaire et Cellulaire, Strasbourg | IGBMC | Research profile

Promoters of ASCL1‐ and NEUROD1‐dependent genes are specific targets of lurbinectedin in SCLC cells | EMBO Molecular Medicine

Active mRNA degradation by EXD2 nuclease elicits recovery of transcription after genotoxic stress | Nature Communications















